[ad_1]
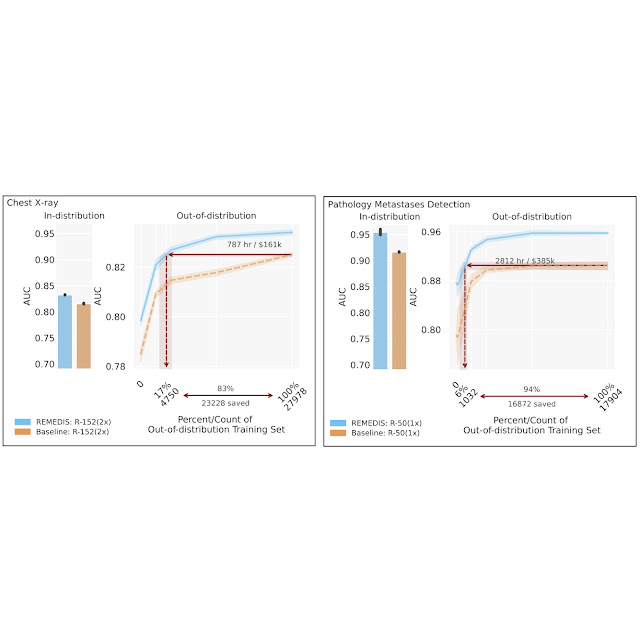
Despite recent progress in the field of medical artificial intelligence (AI), most existing models are narrow, single-task systems that require large amounts of labeled data for training. Moreover, these models cannot be reused in new clinical contexts because they often require site-specific data collection, de-identification and annotation for each new deployment environment, which is time-consuming and expensive. this problem Effective data generalization (the ability of a model to generalize to new settings using minimal new data) remains a key translational challenge for medical machine learning (ML) models and, in turn, prevents their widespread adoption in real-world healthcare settings.
The emergence of foundation models provides an important opportunity to rethink the development of medical AI to make it more effective, safer and fairer. These models are trained using a range of data, often with self-supervised learning. This process leads to generalist models that can be quickly adapted to new tasks and environments with less need for supervised data. With foundational models, it may be possible to use the models safely and effectively in a variety of clinical contexts and settings.
“Robust and Efficient Self-Supervised Medical Imaging” (REMEDIS), Publ Nature is biomedical engineering, we present a unified large-scale self-supervised learning framework for building foundational medical image models. This strategy combines large-scale supervised transfer learning with self-supervised learning and requires minimal task customization. REMEDIS demonstrates a significant improvement in effective data generalization by reducing medical imaging tasks and modalities by 3–100 times to site-specific data for adapting models to new clinical contexts and environments. Based on this, we are pleased to announce the expansion of the Medical AI Research Foundations (hosted by PhysioNet), chest X-ray Foundations public offering in 2022. Medical AI Research Foundations is a collection of open source non-diagnostic models (starting with the REMEDIS models), APIs, and resources to help researchers and developers accelerate medical AI research.
Large-scale self-supervision for medical imaging
REMEDIS uses a combination of natural (non-medical) images and unlabeled medical images to build powerful medical imaging foundation models. His pre-training strategy consists of two stages. The first involves supervised representation training on a large-scale database of labeled natural images (output from Imagenet 21k or JFT) using the Big Transfer (BiT) method.
The second step involves intermediate self-supervised learning that does not require any labels and instead trains the model to learn to represent medical data independent of labels. A specific approach used to represent pre-training and learning is SimCLR. The method works by maximizing the agreement between differently enhanced views of the same training example, through contrast loss, in the hidden layer of a feedforward neural network with the outputs of a multi-layer perceptron (MLP). However, REMEDIS is equally compatible with other contrasting self-supervised learning methods. This training method is used in the healthcare environment as many hospitals acquire raw data (images) as a routine practice. Although processes must be in place to allow these data to be used within models (ie, patient consent prior to data collection, de-identification, etc.), the costly, time-consuming, and complex task of labeling these data can be avoided. Using REMEDIS.
 |
| REMEDIS uses large-scale supervised learning using natural images and self-supervised learning using unlabeled medical data to build robust foundational models for medical imaging. |
Given the limitations of ML model parameters, it is important that our proposed approach works when using both small and large model architecture sizes. To study this in detail, we considered two ResNet architectures with commonly used depth and width multipliers, ResNet-50 (1×) and ResNet-152 (2×), as core encoder networks.
After pre-training, the model was refined using labeled task-specific medical data and evaluated for distributional task performance. In addition, to assess the effective generalization of the data, the model was also optionally specified with a small amount of out-of-distribution (OOD) data.
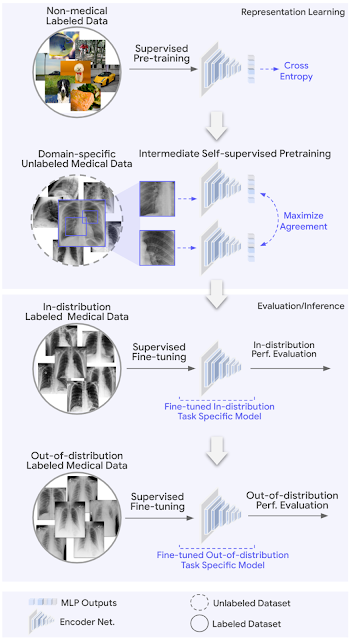 |
| REMEDIS starts with representations that are initialized using large-scale natural image pre-training using the Big Transfer (BiT) method. We then adapt the model to the medical domain using intermediate contrast self-supervised learning without using any labeled medical data. Finally, we tune the model for specific medical imaging tasks. We evaluate the ML model in both in-distribution (ID) and out-of-distribution (OOD) settings to determine the effective generalization of the model to the data. |
Evaluation and results
To evaluate the performance of the REMEDIS model, we run realistic scenarios using retrospective de-identified data across a wide range of medical imaging tasks and modalities, including dermatology, retinal imaging, chest X-ray interpretation, pathology, and mammography. We further introduce the notion of efficient data generalization, which refers to the model’s ability to generalize to new layout distributions with a greatly reduced need for expert annotated data from a new clinical setting. Performance in distributions is measured as (1) zero generalization improvement over OOD parameters (estimating performance on the OOD evaluation set with zero access to training data from the OOD dataset) and (2) significant reduction in the need for annotation. Data from OOD parameters to achieve efficacy equivalent to clinical experts (or threshold for demonstration of clinical benefit). REMEDIS exhibits significantly improved performance in distribution with up to 11.5% improved diagnostic accuracy over a strictly supervised baseline.
Most importantly, our strategy leads to efficient data generalization of medical imaging models, matching strong supervised baselines, resulting in a 3-100-fold reduction in the need for data retraining. Although SimCLR is the primary self-learning approach used in the study, we also show that REMEDIS is compatible with other approaches such as MoCo-V2, RELIC, and Barlow Twins. In addition, the approach works within the dimensions of the model architecture.
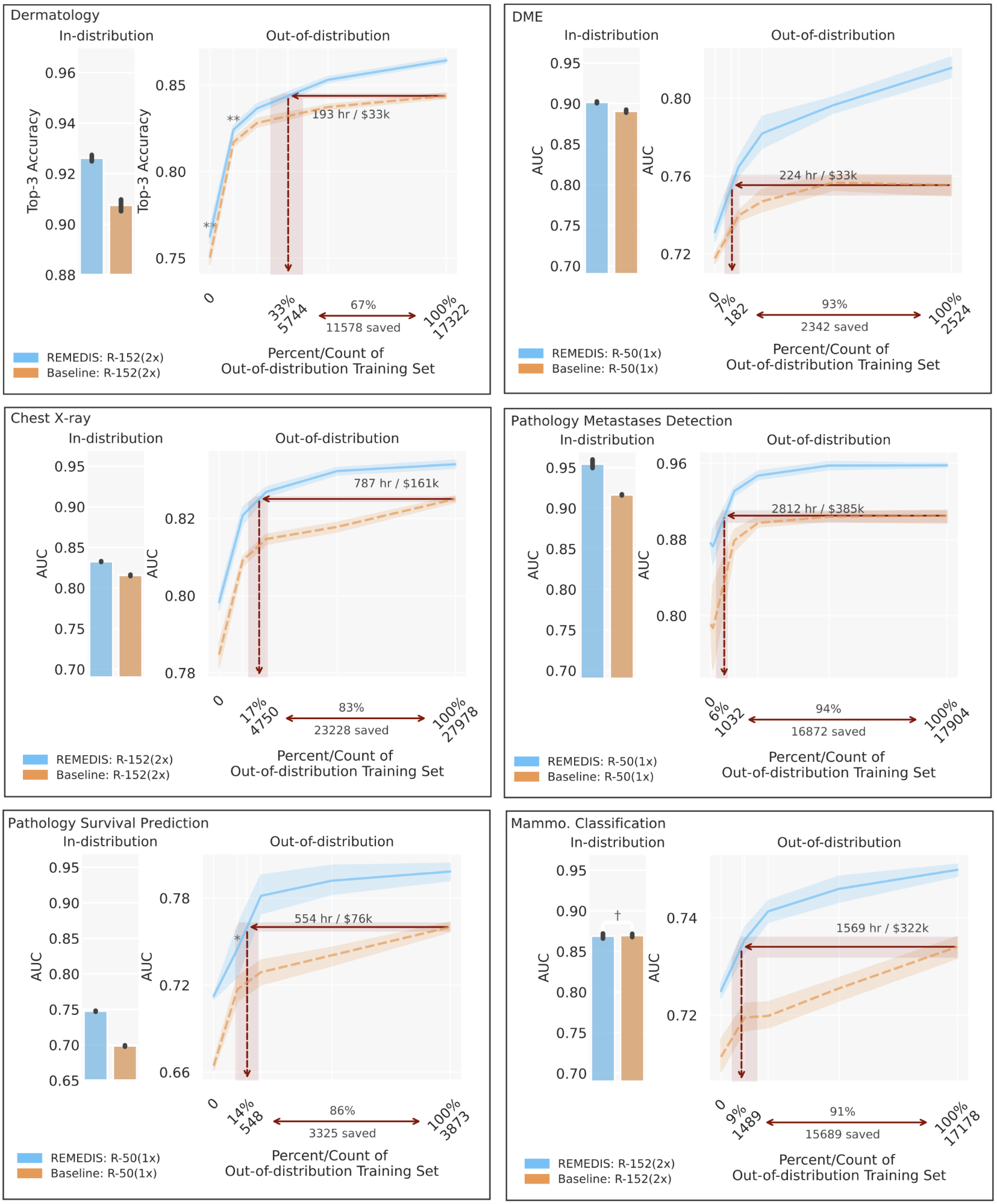 |
| REMEDIS outperformed initial JFT-300M pre-trained supervision for a variety of medical tasks and demonstrated improved data generalization, reducing data requirements by 3-100 times to adapt models to new clinical settings. This could potentially translate into significant reductions in clinic hours through saved annotated data and the development of robust medical imaging systems. |
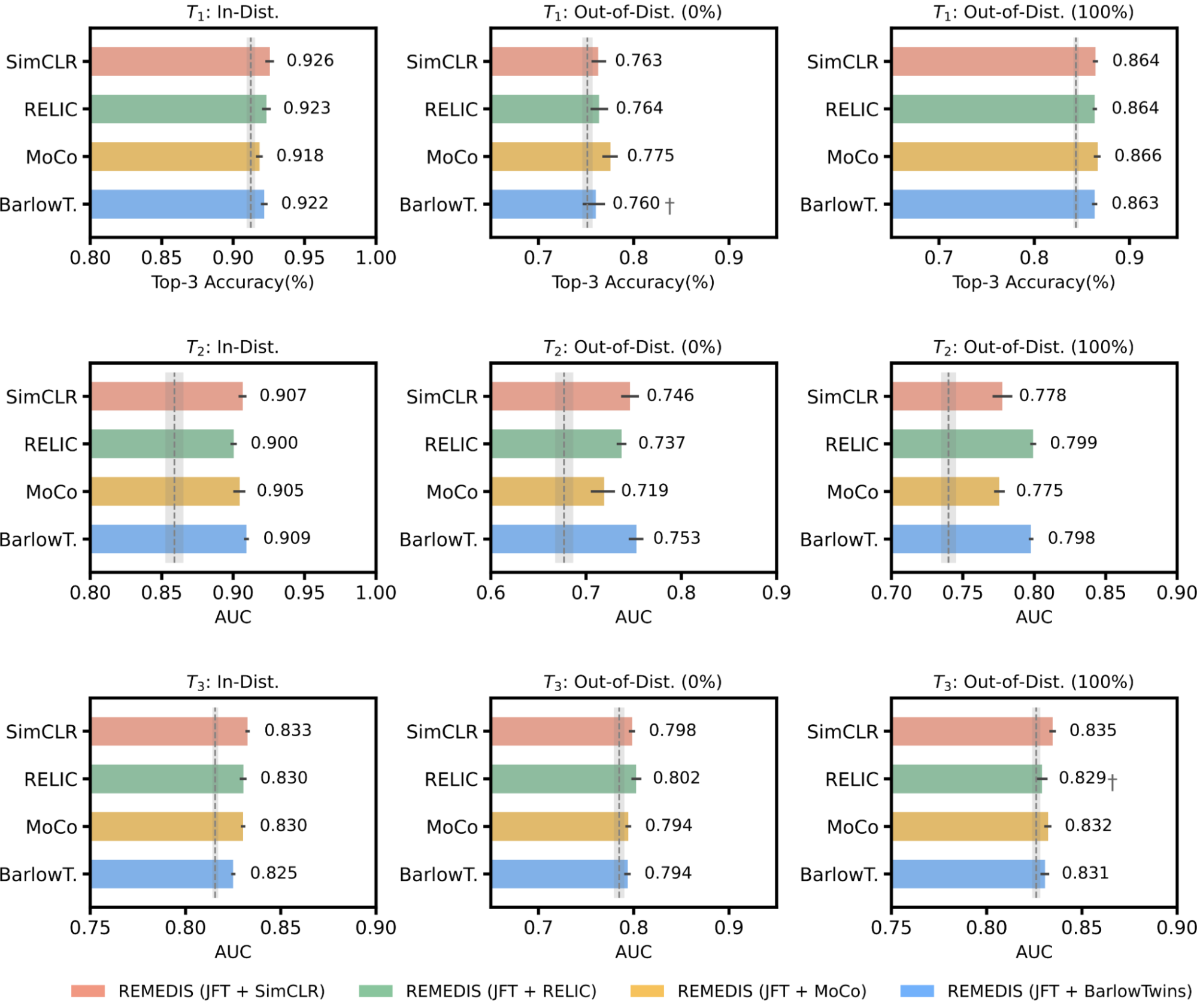 |
| REMEDIS is compatible with MoCo-V2, RELIC, and Barlow Twins as alternative self-supervised learning strategies. All variants of REMEDIS lead to an improvement in the effective generalization of the data compared to the strong supervised baseline for classification of dermatological conditions (T1Classification of diabetic macular edema (T2), and chest X-ray status classification (T3). The gray shaded area indicates the performance of the strongly supervised baseline that was pretrained on the JFT. |
Medical AI Research Foundations
Building on REMEDIS, we are excited to announce the Medical AI Research Funds, an expansion of the Chest X-ray Funds public offering in 2022. Medical AI Research Foundations is an open source repository of medical foundation models hosted by PhysioNet. This extends the previous API-based approach to also include non-diagnostic models to help researchers and developers accelerate medical AI research. We believe that the release of REMEDIS and the Medical AI Research Funds is a step towards creating medical models that can generalize across healthcare settings and tasks.
We are seeding medical AI research funds with REMEDIS models for chest x-ray and pathology (linked code). While the existing chest X-ray foundation approach focuses on providing frozen builds for application-specific fine-tuning from a model trained on several large private datasets, REMEDIS models (trained on public datasets) allow users to fine-tune them end-to-end for use and running on local devices. We recommend that users try different approaches based on their unique needs for their desired application. We expect to add more models and resources for training medical foundation models, such as datasets and benchmarks, in the future. We also welcome the medical AI research community to contribute to this.
conclusion
These results suggest that REMEDIS has the potential to significantly accelerate the development of ML systems for medical imaging that can maintain their robust performance when deployed in a variety of changing contexts. We believe this is an important step forward for artificial intelligence in medical imaging to have a broad impact. In addition to the experimental results presented, the approach and insights described here have been integrated into several Google medical imaging research projects, such as dermatology, mammography, and radiology, among others. We use a similar self-supervised learning approach in our non-representative foundation model efforts such as Med-PaLM and Med-PaLM 2.
With REMEDIS, we have demonstrated the potential of foundational models for medical imaging applications. Such models have interesting possibilities in medical applications with the ability to learn multimodal representations. The practice of medicine is inherently multimodal and integrates information from images, electronic health records, sensors, wearables, genomics, and more. We believe that ML systems that use this data at scale using self-supervised learning with careful consideration of privacy, security, fairness and ethics will help lay the foundation for the next generation of learning healthcare systems that will bring world-class healthcare to everyone at scale.
Acknowledgments
This work involved a broad collaborative effort from a multidisciplinary team of researchers, software engineers, clinicians, and cross-functional contributors across Google Health AI and Google Brain. In particular, we would like to thank our co-first author Jan Freiberg and our lead senior authors Vivek Natarajan, Alan Karthikesalingam, Mohammad Norouzi and Neil Houlsby for their invaluable input and support on these projects. We also thank Lauren Wiener, Sami Lachgar, Yun Liu, and Karan Singhal for their feedback on this post, and Tom Small for his support in creating the visuals. Finally, we also thank the PhysioNet team for their support in hosting the Medical AI Research Funds. Users with questions can contact medical-ai-research-foundations at google.com.
[ad_2]
Source link

