[ad_1]
An interesting (and beneficial, despite an otherwise less-than-ideal situation) side effect of the COVID-19 pandemic has been that many organizations, commercial and academic, are looking for ways they can work together to eradicate it. disease. Many collaborations involve sharing datasets, the best known of which is the COVID-19 Open Research Dataset (CORD-19), a collection of 47,000 (and growing) scientific papers on COVID-19. Some others offer CORD-19 databases processed by their pipelines or hosted using their products (for example, a graph database or search engine). Some hold workshops and share their experiences with the rest of the world. At Elsevier, we have a data science team of more than 225 employees who look at the problem from different disciplines and angles and work to find solutions to solve the crisis. The LinkedIn article Elsevier’s Models for COVID19 Biomolecular Mechanisms describes some of the contributions that have resulted from this team’s work using one of our tools, and hopefully more to come soon. In addition, about 46% of the papers in the CORD-19 dataset come from Elsevier, and we are looking at ways to make it more accessible.
In the spirit of learning all I could about COVID-19, I attended the one-day COVID-19 and AI: Virtual Conference organized by the Stanford Human-AI (HAI) group. One of the speakers was Professor Nigam Shah, who spoke about his medical center’s data science response to the pandemic and described the types of data science models that can inform policy to combat the virus. Additionally, he also wrote this Medium post about Symptom Profiling of SARS-Cov-2 Screened Patients where he used the same chart for his unified model, which caught my eye. Hat tip to my colleague Helena Deus for finding and posting a link to the article on our internal Slack channel.
In any case, the Medium post describes a text-processing pipeline developed by Professor Nigam’s group to extract clinical observations from notes written by emergency department providers at Stanford Health Care while screening patients for COVID-19. The pipeline is built using similar rules based on the NegEx algorithm, among other things, and Snorkel to train models that recognize these observations in text using these noisy rules. The frequencies of these observations were then tallied and probabilities were calculated, eventually leading to the Excel spreadsheets that Professor Niga and his team were kind enough to share with the world.
For this data set, 895 patients were considered, of which 64 tested positive for SARS-Cov-2 (new name is COVID-19) and 831 tested negative. So, at this point in time, the prevalence of COVID-19 in the cohort (and by extension, possibly in the wider community) was 7.2%. Observations considered in the model were those that occurred at least 10 times in all patient records.
What can we do with this data? My first thought was a symptom test, which would calculate the probability that a particular patient is positive given one or more observations (or symptoms, although I’m using the term a bit loosely, there are quite a few observations that aren’t symptoms). For example, if we wanted to calculate the probability of a patient testing positive given that the patient only has a cough and no other symptoms, we would denote this as P(D=True|S).0= True, S1=false, …, s49= false).
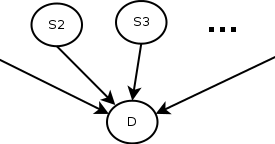
Of course, this depends on the simplistic (and most likely incorrect) assumption that the observations are independent, that is, the fact that the patient has a cough is independent of the fact that he has a sore throat. Also, another thing to remember is that the symptom checker’s predictions will depend on the correct value of the current prevalence rate of the disease. The 7.2% value we have is only correct for the time and place the data was collected, so it will need to be updated accordingly if we are to use the check, even with all its limitations. Here is a schematic of the model.
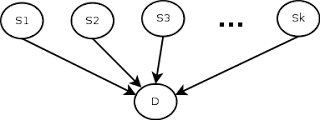
In terms of implementation, I initially considered a Bayesian network, using SQL tables to model it, as taught by Professor Gautam Shroff in his recently released Web Intelligence and Big Data course on Coursera (here’s a quick note on how to use SQL tables to model Bayesian networks, (because the technique, although it’s very cool, doesn’t seem mainstream), but I realized (thanks to a mathematical StackExchange discussion about conditional probability expressions considering many independent events) that the formulation could be much simpler as shown. below, so I used that instead.
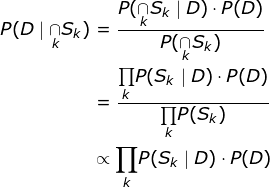
The idea behind using the proportionality relationship is to normalize the numerator by computing P(D=True|∩S).K).P(D=True) and P(D=False|∩SK).P(D=false) and divide by the sum to get the probability of a positive test given a set of symptoms. Once this was done, it led to some additional interesting questions. First, what happens to the probability as we add more and more symptoms? Second, what happens to the probability with different spreading rates? Finally, what is the “symptom profile” for a typical COVID-19 patient based on the data? The answers to these questions and the code to get these answers can be found in my Github Gist here .
I’ve said it before, and given that the pandemic may have people grasping at straws, I’m going to say it again. This is only a model and most likely imperfect. Findings from such models are not a substitute for medical advice. I know most of you already realize this, but just in case, please do not use the conclusions of this model to make any real-life decisions without independent verification.
[ad_2]
Source link